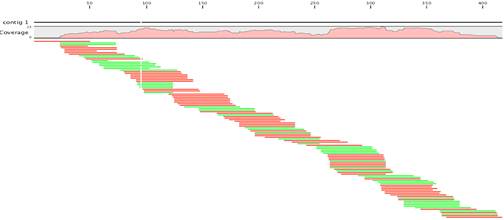
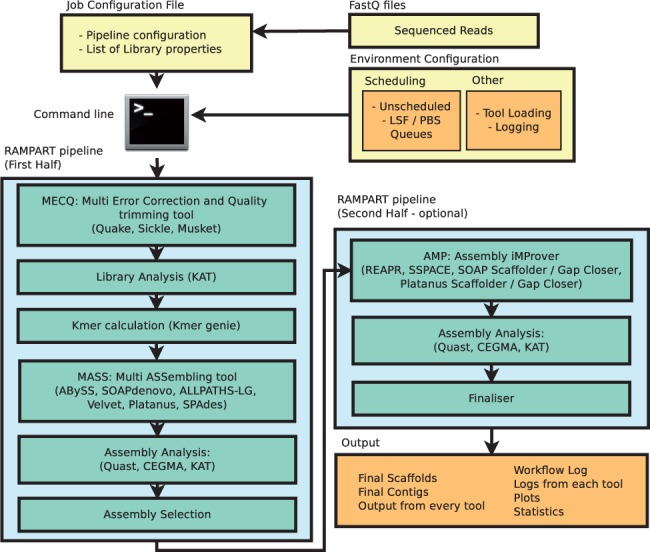
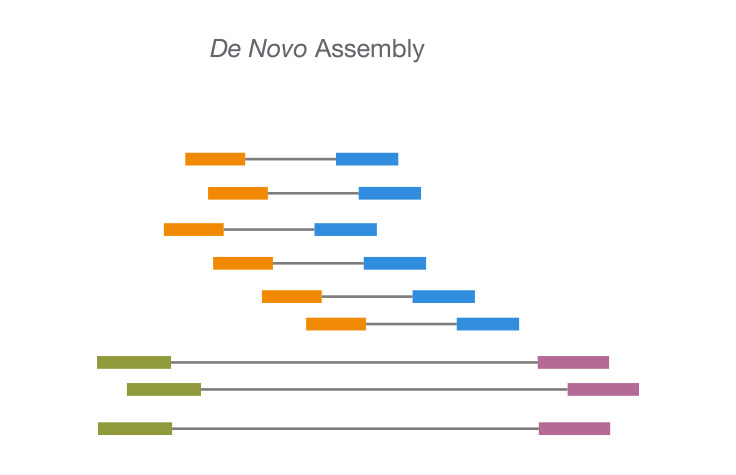
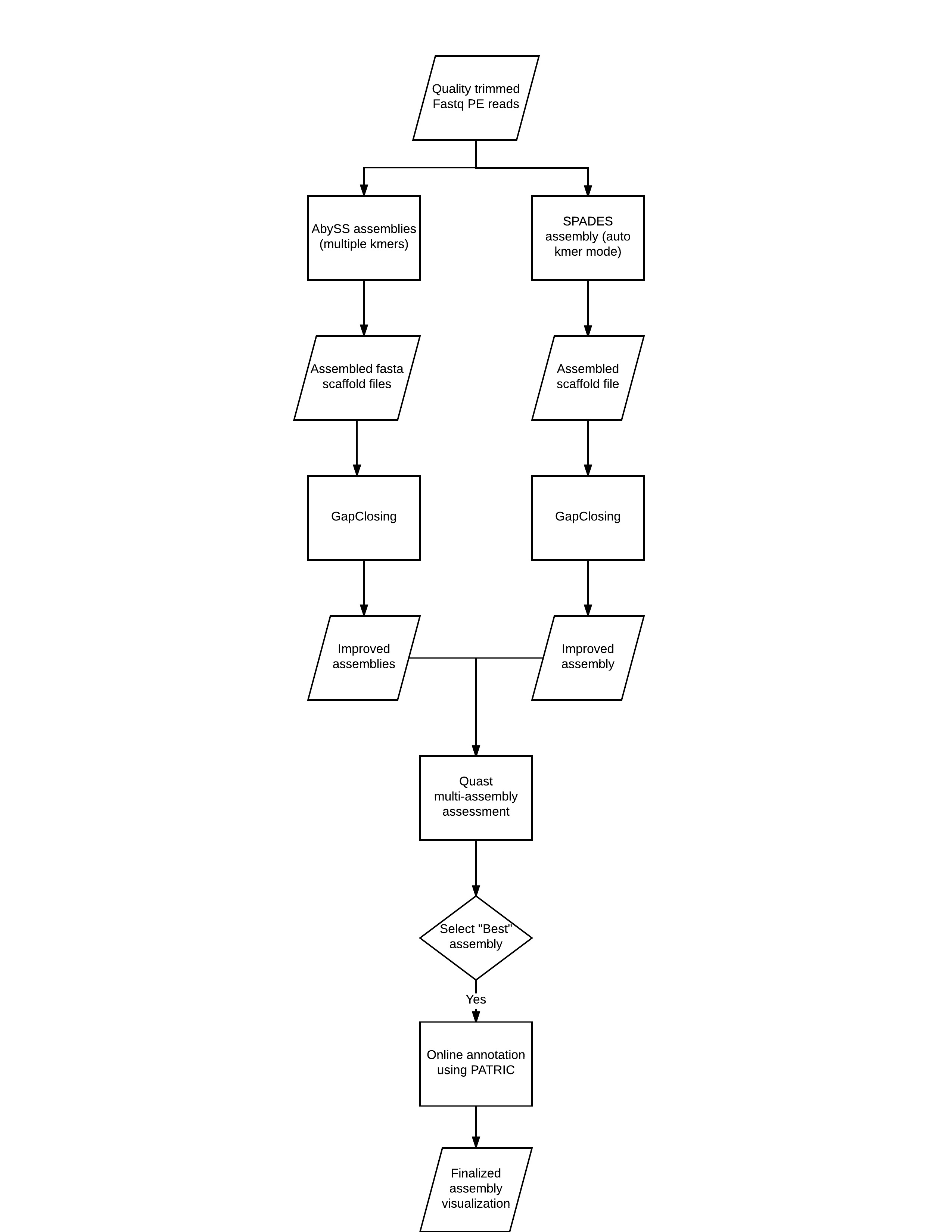
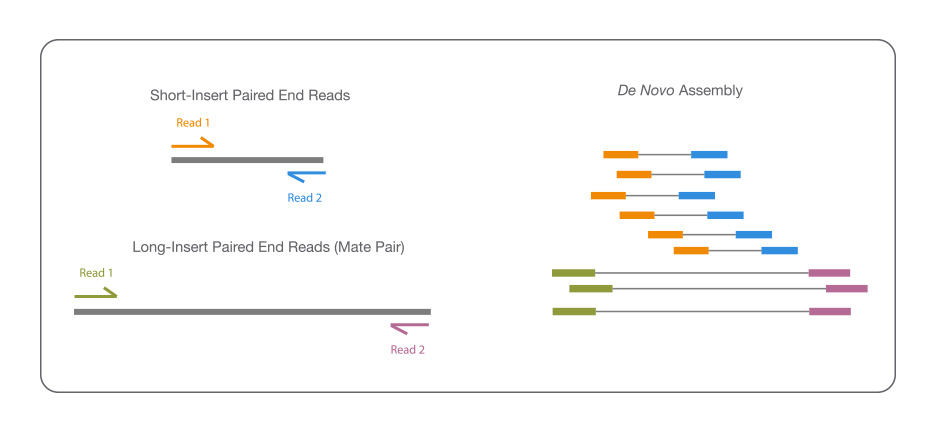
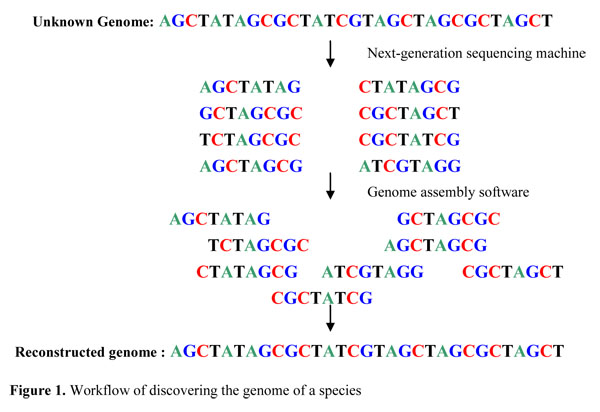
Rapid hybrid de novo assembly of a microbial genome using only short reads: Corynebacterium pseudotuberculosis I19 as a case study - ScienceDirect
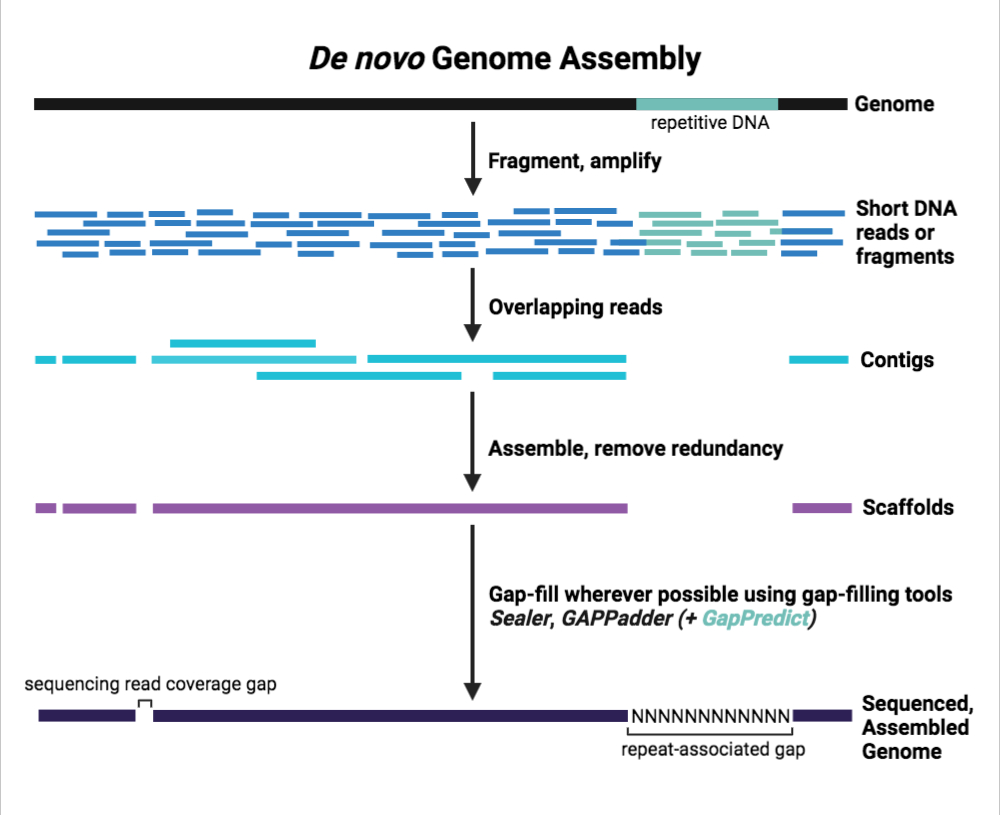
Filling in the gaps: GapPredict can complement repertoire of tools used to resolve missing DNA sequences in genome assemblies | Genome Sciences Centre
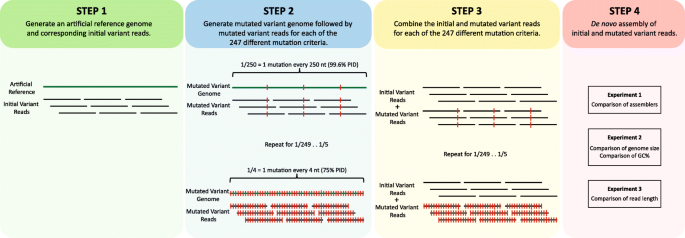
The effect of variant interference on de novo assembly for viral deep sequencing | BMC Genomics | Full Text
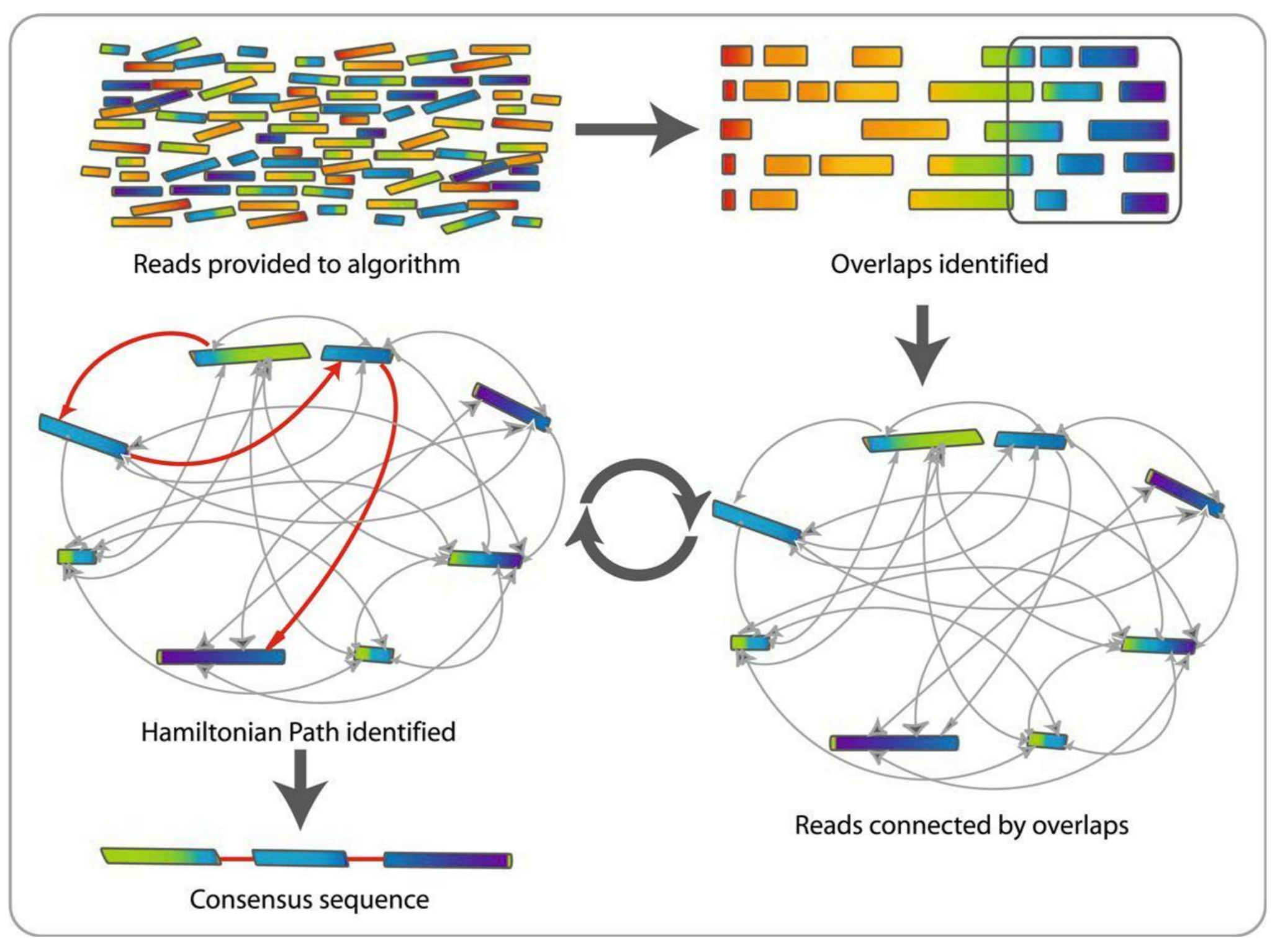
Slide Deck: Slide Deck: Hands-on: Unicycler assembly of SARS-CoV-2 genome with preprocessing to remove human genome reads / Assembly

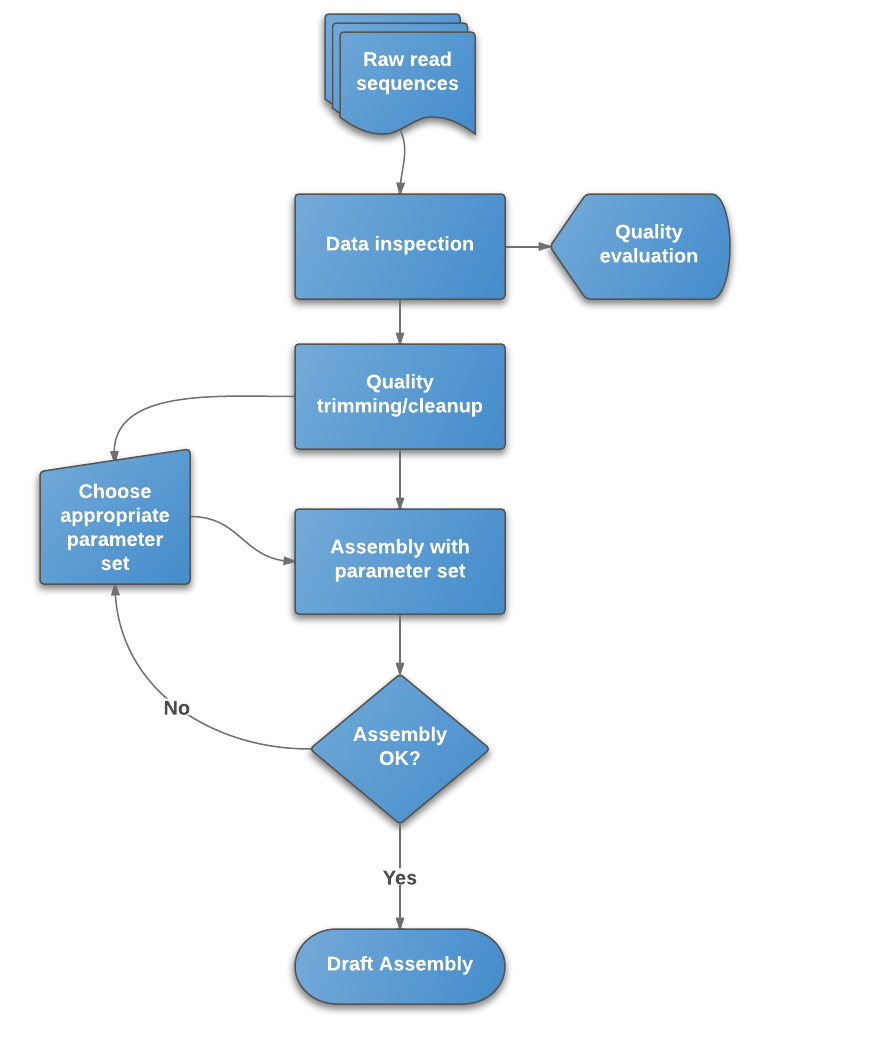
Whole Genome de novo Assembly |Supporting de novo assembly training |Course |WGS sequencing | data analysis| Workshop

Comparison of reference assembly and de novo assembly. A) Reference... | Download Scientific Diagram

Sequencing from scratch: reference genomes and de novo sequence assembly – HudsonAlpha Institute for Biotechnology
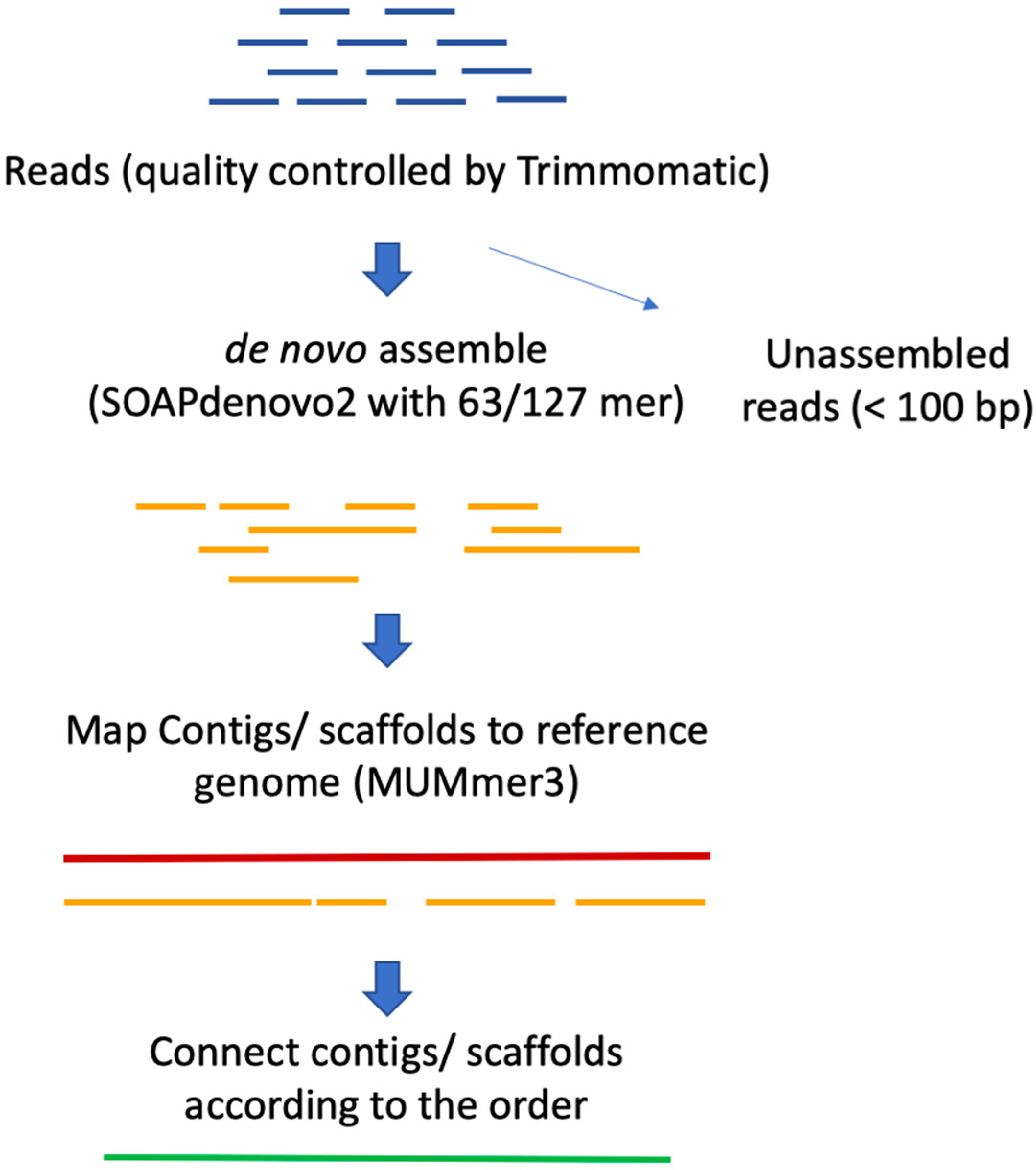
Plants | Free Full-Text | Reference-Guided De Novo Genome Assembly to Dissect a QTL Region for Submergence Tolerance Derived from Ciherang-Sub1

High-throughput pipeline for the de novo viral genome assembly and identification of minority variants from Next-Generation Sequencing of residual diagnostic samples | bioRxiv